The APEER platform allows creating modules and workflows using basically any programming language due to its underlying Docker(TM) technology. Nevertheless, Python seems to be the favorite choice for most of the time for various reasons:
- Simple enough to be used by researchers with scripting experience
- Powerful enough to create amazing computational tools
- A huge and very open community and the whole ecosystem behind it
- Probably the most popular language when it comes to topics like Machine Learning
The topic or question of what is the "best" image data format for microscopy is a very interesting and also quite difficult question. There are no easy answers and there is no right or wrong here.
Our goal was a simple python-based tool to read and write OME-TIFF
Since the APEER platform tries to constantly provide solutions for our users our team decided to support the currently most popular image data format for microscopy image data, which cleary is OME-TIFF (despite its known limitations). Therefore, we explored easy and simple ways to read OME-TIFF for the most common use cases. We just wanted a simple python-based tool to read and write OME-TIFF without the need to include JAVA etc. into the modules. We reused parts of the existing python ecossystem, especially python-bioformats and TIFF-file, added some extra code and created a basic PyPi package.
This package can be easily inclued in every APEER module but it can also be used inside our python application or within jupyter notebook.
How to use the OME-TIFF-Library
In order to make things a bit easier we create a little helper script for you.
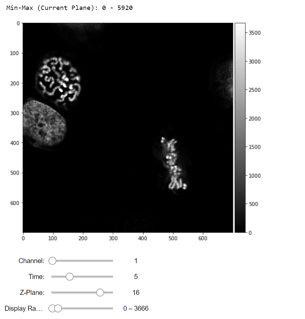
Reading the OME-TIFF stack as an NumPy Array
To understand how we did this, please refer to our helper script.
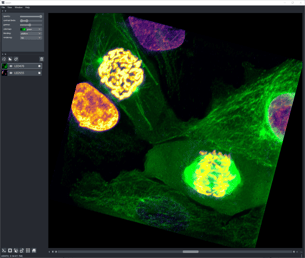
Here we use the Napari viewer to visualize the complete OME-TIFF stack, which is represented by a multi-dimensional NumPy Array.